The global rise in antibiotic resistance in recent years has given rise to a proliferation of multidrug-resistant (MDR) bacteria capable of causing life-threatening infections. The challenge is particularly pronounced in developing countries such as Bangladesh, where antibiotics are overprescribed and often available from pharmacies without prescription.1
Many Bangladeshis do not have access to clean drinking water, leading to a high prevalence of waterborne diseases such as Salmonella infections.2 Widespread use of antibiotics over the years has caused MDR Salmonella strains to proliferate, rendering several commonly used antibiotics ineffective.3,4
Salmonella is responsible for causing salmonellosis, typhoid fever, gastroenteritis, and bacteremia.5 More than 2700 serotypes of Salmonella have been identified.6 Enteric infections caused by Salmonella are responsible for causing approximately 4500 deaths each year in Bangladesh.7 Chloramphenicol was used as a first-line therapy for the treatment of Salmonella infections till growing resistance limited its use.4-6 A study in Chattogram, Bangladesh, found that typhoid patients were not responsive to ciprofloxacin, a second-line therapy.4 Antibiotic resistance generally develops in Salmonella strains through mutations in chromosomal genes and plasmids. The transmission of the genes is promoted significantly by horizontal gene transfer.8
A significant reason for the rapid emergence of MDR strains is the excessive use of antibiotics for prevention, treatment, and metaphylaxis.9 A study in southern rural Bangladesh found that around 44% of doctors prescribe antibiotics without a clinical diagnosis.4 Another reason is poor patient adherence, many of whom do not understand the impact of incomplete doses.10 Also, the uncontrolled use of prophylactic antibiotics in farm animals vastly contributes to antibiotic resistance.11
The lack of comprehensive data on antibiotic resistance patterns in Bangladesh has led to suboptimal therapeutic choices for treating infections.4 To ensure optimal treatment of Salmonella infections in Bangladesh, it is necessary to determine the prevalence of resistance against the commonly used antibiotics. Therefore, this study aimed to identify the prevalence of resistance of Salmonella spp. against commonly used antibiotics in the city of Dhaka.
Methods
Clinical samples were collected from different diagnostics centers across Dhaka from February to July 2023. A total of 150 samples (135 urine, 13 sputum, and two tracheal aspirates), were collected in a sterile container, transported to the laboratory, and stored at 4 °C for further analysis.
Salmonella spp. was isolated from the clinical samples by streak plate method using solidified Xylose Lysine Deoxycholate agar and Salmonella-Shigella agar media. A single isolate per sample was subjected to antibiotic susceptibility analysis to avoid redundancy or incidence of duplication. After incubation at 37 °C, the bacterial growth was assessed. Salmonella species developed red colonies with a black center on Xylose Lysine Deoxycholate agar plate. In Salmonella-Shigella agar, non-lactose fermenting, colorless colonies with black centers, were identified as Salmonella isolates. The selected colonies were characterized by Gram staining. For standard biochemical assays, the Indole test, Triple Sugar Iron test, Urease test, Methyl Red/Voges-Proskauer test, Oxidase test, and Citrate Utilization test were carried out.12,13
The antibiotic susceptibility of the isolates was tested using the Kirby-Bauer Disc Diffusion Method.13 We used 12 commercially available antibiotics which belonged to nine different classes as follows: (1) penicillins: penicillin G (30 µg) and piperacillin (100 µg), (2) cephalosporins: cefuroxime (30 µg), (3) tetracyclines: tetracycline (30 µg), (4) macrolides: azithromycin (30 µg) and clindamycin (10 µg), (5) fluoroquinolones: norfloxacin (10 µg), (6) sulphonamides: cotrimoxazole (25 µg), (7) aminoglycosides: gentamicin (10 µg), (8) chloramphenicol (30 µg), and (9) carbapenem: imipenem (10 µg) and meropenem (10 µg). Based on action mechanisms, the antibiotics used in this study belonged to three major groups: (a) bacterial cell wall synthesis inhibitors (piperacillin, penicillin G, cefuroxime, imipenem, and meropenem); (b) bacterial protein synthesis inhibitors (tetracycline, gentamicin, clindamycin, chloramphenicol, and azithromycin); and (c) bacterial nucleic acid synthesis inhibitors (norfloxacin and cotrimoxazole).
The culture samples were spread on Mueller-Hinton Agar media and the antibiotic discs were placed. After incubation at 37 °C, the diameter of the clear zones around the antibiotic discs was measured and the isolates were categorized as sensitive, intermediate, or resistant according to the Clinical Laboratory Standard Institute guidelines.13,14 Standard bacterial strains were used as control to validate the results.
The isolates showing resistance to ≥ 3 classes of antibiotics were considered MDR. The isolates exhibiting susceptibility to only 1–2 classes of antibiotics were considered extensively drug-resistant (XDR). The multiple antibiotic resistance (MAR) index was determined using the formula, x/y, where ‘x’ is the number of isolates resistant to antibiotics and ‘y’ is the total number of antibiotics used on each isolate.
Results
From a total of 150 clinical samples, 100 Salmonella isolates were obtained, representing a prevalence of 66.7%. The sources were urine (90.0%), sputum (9.0%), and tracheal aspirate (1.0%).
Gram staining test on the Salmonella isolates revealed rod-shaped, pink-colored colonies forming a chain. All the isolates showed a positive citrate test evidenced by the media turning an alkaline blue. A positive triple sugar iron test was confirmed by a reddish-black slant and yellow butt, indicating production of hydrogen sulfide gas. The indole test was negative for the isolates, indicated by the absence of a red-violet ring on the surface of the liquid culture media. The isolates showed a negative result in the Voges-Proskauer test and a positive result in the Methyl Red test, as evidenced by the development of a brownish-yellow color and a cherry red color, respectively. All the isolates showed positive results for motility tests. The catalase, Oxidase, and Urease tests for the isolates yielded negative results.
The susceptibility of all 100 Salmonella isolates was evaluated against 12 different antibiotics and the diameters of the zones of inhibition were measured [Table 1]. The isolates were classified as resistant, intermediate, or sensitive to a particular antibiotic using a zone diameter interpretive standards chart. The isolates showed varying resistance against the 12 antibiotics [Table 2 and Figure 1].
Table 1: Zones of inhibition (mm) against different antibiotics for categorizing Salmonella isolates as resistant, intermediate, and sensitive (N = 100).
|
Tetracycline
|
9.4 ± 0.3
|
13.3 ± 0.3
|
18.3 ± 0.4
|
|
Penicillin G
|
9.7± 0.4
|
-
|
29.4 ± 0.5
|
|
Piperacillin
|
9.7 ± 0.4
|
-
|
16.0 ± 0.3
|
|
Clindamycin
|
10.0 ± 0.4
|
15.1 ± 0.3
|
21.7 ± 0.4
|
|
Meropenem
|
10.5 ± 0.5
|
17.2 ± 0.6
|
23.6 ± 0.5
|
|
Imipenem
|
9.9 ± 0.4
|
14.2 ± 0.4
|
19.9 ± 0.4
|
|
Azithromycin
|
10.2 ± 0.4
|
13.9 ± 0.3
|
18.6 ± 0.3
|
|
Cotrimoxazole
|
9.2 ± 0.5
|
13.1 ± 0.6
|
19.0 ± 0.6
|
|
Gentamicin
|
10.5 ± 0.5
|
13.5 ± 0.5
|
18.4 ± 0.5
|
|
Chloramphenicol
|
10.6 ± 0.5
|
11.8 ± 0.5
|
22.3 ± 0.5
|
|
Norfloxacin
|
11.4 ± 0.6
|
14.8 ± 0.8
|
20.8 ± 0.5
|
Table 2: Resistance exhibited by Salmonella isolates against 12 different antibiotics (N = 100).
|
Tetracycline
|
50
|
16
|
34
|
|
Penicillin G
|
96
|
0
|
4
|
|
Piperacillin
|
61
|
0
|
39
|
|
Clindamycin
|
71
|
12
|
17
|
|
Meropenem
|
43
|
8
|
49
|
|
Imipenem
|
33
|
11
|
56
|
|
Azithromycin
|
43
|
17
|
40
|
|
Cotrimoxazole
|
68
|
9
|
23
|
|
Gentamicin
|
32
|
8
|
60
|
|
Chloramphenicol
|
50
|
14
|
36
|
|
Norfloxacin
|
27
|
13
|
60
|
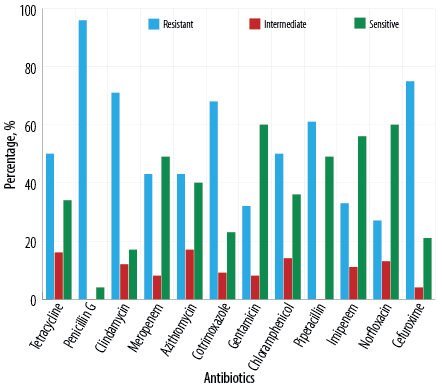 Figure 1: Antibiotic resistance pattern among Salmonella isolates (N = 100).
Figure 1: Antibiotic resistance pattern among Salmonella isolates (N = 100).
As shown in Table 2, the highest prevalence of resistance was observed against penicillin G (96.0%), followed by cefuroxime (75.0%), clindamycin (71.0%), cotrimoxazole (68.0%), and piperacillin (61.0%). Against tetracycline and chloramphenicol, the isolates showed 50.0% resistance each.
A Significantly high degree of resistance was also observed against meropenem (43.0%), azithromycin (43.0%), imipenem (33.0%), and gentamicin (32.0%). The lowest resistance was observed against norfloxacin (27.0%).
The majority of the isolates were sensitive to three antibiotics: gentamicin (60.0%), norfloxacin (60.0%), and imipenem (56.0%). The other antibiotics triggered sensitivities < 50%. Among the antibiotics, penicillin G was the least effective with only 4.0% sensitivity and 0.0% intermediate susceptibility. Seventeen percent of the isolates showed intermediate susceptibility to azithromycin.
The cell wall synthesis inhibitor antibiotics included penicillin G and piperacillin from the penicillin group, imipenem and meropenem from the carbapenems, and cefuroxime from the cephalosporins. Among these five antibiotics, Salmonella isolates exhibited the highest resistance against penicillin G (31.0%) and the lowest resistance against imipenem (11.0%) [Figure 2]. Among the cell wall inhibitory agents, the carbapenem group had the highest sensitivity (62.0%).
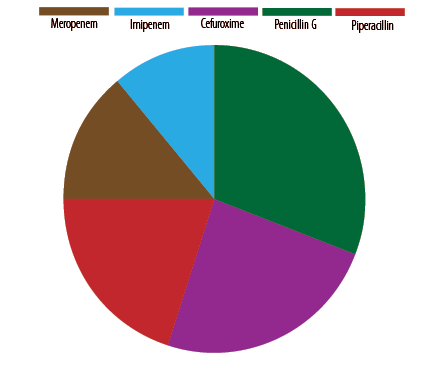 Figure 2: Resistance pattern among Salmonella isolates based on cell wall synthesis inhibitors
Figure 2: Resistance pattern among Salmonella isolates based on cell wall synthesis inhibitors
(N = 100).
We used five bacterial protein synthesis inhibitors: azithromycin, chloramphenicol, clindamycin, gentamicin, and tetracycline. The highest resistance was observed against clindamycin (29.0%), followed by tetracycline (21.0%), chloramphenicol (21.0%), azithromycin (18.0%), and gentamicin (11.0%) [Figure 3]. On the other hand, the Salmonella isolates showed the highest sensitivity towards gentamicin (32.0%), followed by azithromycin (22.0%), and chloramphenicol (19.0%) among the bacterial protein synthesis inhibitors.
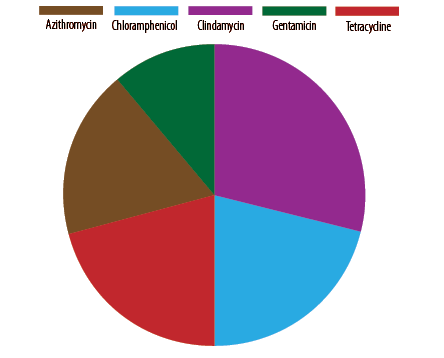 Figure 3: Resistance pattern among the Salmonella isolates based on protein synthesis inhibitors
Figure 3: Resistance pattern among the Salmonella isolates based on protein synthesis inhibitors
(N = 100).
Between the two nucleic acid synthesis inhibitor antibiotics, the resistance was higher against norfloxacin (72.0%) than against cotrimoxazole (28.0%).
Among the Salmonella isolates from urine samples (n = 90), resistance was highest against penicillin G (96.0%), followed by cefuroxime (77.0%), clindamycin (70.0%), and cotrimoxazole (66.0%). Resistance to piperacillin, tetracycline, chloramphenicol, and azithromycin ranged 40–60%. Carbapenem antibiotics also encountered significant resistance; imipenem (32.0%) and meropenem (38.0%). Gentamicin was effective against 65.0% urine isolates and 78.0% sputum isolates. Among the 13 sputum samples, nine Salmonella strains were isolated. Their resistance against various antibiotics were: penicillin G (89.0%), meropenem (89.0%), clindamycin (78.0%), cotrimoxazole (78.0%), tetracycline (52.0%), imipenem (45.0%), piperacillin (55.0%), and chloramphenicol (48.0%).
As shown in Figure 4, 98.0% of isolates showed a MAR index value of ≥ 0.2, while 90.0% exhibited MAR index value of ≥ 0.3, and 65.0% had MAR index value of ≥ 0.5. The results revealed a high prevalence of multidrug resistance as 98.0% of the isolates showed MDR, while 23.0% were resistant to five classes of antibiotics [Figure 5]. As shown in Figure 5, 18.0% of the isolates showed resistance towards six classes of antibiotics, and 16.0% of the isolates exhibited resistance against four classes of antibiotics. Two of the isolates were resistant to all the nine classes of antibiotics used in this study. Moreover, in the current study, 20.0% of the isolates showed XDR.
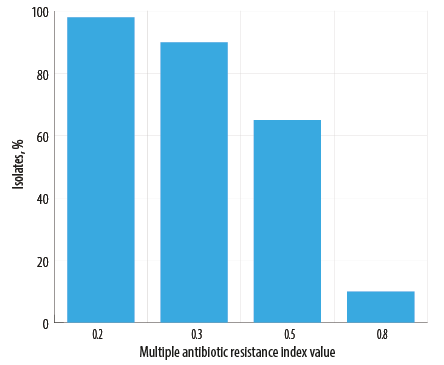 Figure 4: Percentage of isolates with varying Multiple Antibiotic Resistance index values
Figure 4: Percentage of isolates with varying Multiple Antibiotic Resistance index values
(N = 100).
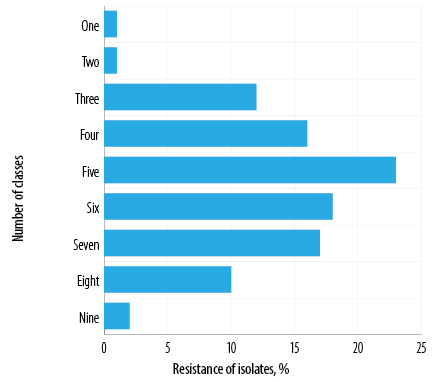 Figure 5: Resistance of Salmonella isolates against multiple classes of antibiotics (N = 100).
Figure 5: Resistance of Salmonella isolates against multiple classes of antibiotics (N = 100).
Discussion
This study was conducted to identify the antibiotic resistance pattern among Salmonella spp. isolated from clinical samples in Dhaka. Nearly all (98.0%) of the 100 Salmonella isolates were MDR, showing resistance against at least three different classes of antibiotics. The rising incidence of antimicrobial resistance caused by MDR pathogens raises serious concerns regarding the therapeutic usability of the available antibiotics in the near future.15
Seventeen percent of the isolates were resistant to seven classes of antibiotics, 18.0% to six classes of antibiotics, and 23.0% to five classes of antibiotics. The results revealed that the most effective antibiotics were gentamicin and norfloxacin, against which 60.0% of the isolates showed sensitivity. Broad-spectrum antibiotics like imipenem or meropenem are increasingly prescribed due to the rising incidence of antimicrobial resistance, though they are reported to be more damaging to human microbiota. In a study carried out in the USA, the KPC-2 β- lactamase-producing strain of Salmonella was found to be resistant to most beta-lactam antibiotics, including oxyimino-cephalosporins and imipenem.16
The current study also revealed the significant prevalence of resistance to carbapenems, which is similar to a finding in Pakistan,17 which also reported high (48%) resistance of Salmonella isolates to meropenem, a carbapenem class antibiotic. The diminishing efficacy of carbapenems against MDR pathogens indicates the urgent need for new alternatives.18
In the current study, 43.0% of the isolates were resistant to azithromycin, similar to a previous study in Bangladesh by Uddin et al,19 which found 40% prevalence of resistance. However, regarding tetracyclines, the resistance found in the current study (50.0%) was nearly double of the 28% reported by Uddin et al,19 An Iranian study in 2016 reported 100% resistance against tetracyclines among Salmonella spp.20 The variation in the resistance could be attributed to the regional differences in disease patterns and MDR pathogens.21
Different studies have reported varying degrees of resistance against chloramphenicol among clinical Salmonella isolates across different countries such as 100% in Saudi Arabia,22 44% in Burkina Faso,23 50% in Kenya,24 and 60% in Pakistan.25 In our study, 50.0% of the isolates showed resistance to chloramphenicol. A previous study (2018) in Bangladesh had reported 32.5% of chloramphenicol resistance, suggesting a significant increase in resistance over a relatively short period.26
In the 1970s, the rise of MDR Salmonella strains against chloramphenicol and others triggered a shift to fluoroquinolones as the preferred therapy in the South and South East Asia.27 Within decades, MDR strains proliferated against fluoroquinolones as shown by the 2009–2011 Salmonella infection data from Pakistan,28 which showed 91.7% resistance to ciprofloxacin, a fluoroquinolone derivative. A subsequent Pakistani study found that the resistance of ciprofloxacin rose marginally to 95%.17 Southeast Asia also reported a significant rise in resistance to fluoroquinolones.29 A study in India focused on the long-term genomic profiling of the resistant genes in Salmonella typhi reported 97% resistance against ciprofloxacin.30 However, other studies reported a much lower prevalence of resistance to fluoroquinolones. From Bangladesh, Mannan et al,31 and Habib et al,32 reported 51% and 20% ciprofloxacin-resistant isolates, respectively. A study conducted in China documented that 20% of clinical Salmonella isolates were resistant to ciprofloxacin.33 In the current study also, only 27.0% of the isolates showed resistance to norfloxacin, a fluoroquinolone molecule.
Against cotrimoxazole (a sulfonamide drug), the Salmonella isolates in the current study exhibited a high resistance of 68.0%. An indication that cotrimoxazole-resistant strains have been proliferating over the years was revealed by a study from Pakistan,34 which reported a resistance prevalence of only 38.5%. A recent Pakistani study reported 66.7% resistance,17 similar to ours. Meanwhile, a study from Bangladesh found 48.4% resistance,26 against cotrimoxazole.
In developing countries, the evolution of MDR bacteria is a significant public health concern.1,10 The problem is mainly related to overusing antibiotics without proper diagnosis.4,13 In the current study, 65.0% of the isolates had MAR index values of ≥ 0.5, indicating a high prevalence of resistance in the samples from Dhaka. Moreover, 20.0% of our isolates showed XDR. However, a large South Asian survey of 8705 enteric fever cases during 2016–2019 reported no XDR isolates from Bangladesh.35 A study in Lahore, Pakistan by Zahid et al,25 reported the presence of 67% MDR along with 34% XDR among Salmonella isolates from clinical samples. More studies from Pakistan revealed strong XDR presence including 35% by Klemm et al,36 and 41.6% by Ambreen et al,37 High prevalence of XDR Salmonella isolates have also been reported from more distant developing countries, such as a 33.3% prevalence in Egypt in 2021.38
Our results demonstrate an alarming rise in the prevalence of MDR and XDR pathogens in Bangladesh, which could be attributed to the irrational use of antimicrobials combined with inadequate AMR surveillance.13 In resource-limited countries such as Bangladesh, the habit of self-medication among patients, and the indiscriminate use of antibiotics in agriculture and farming are a major contributor to the emergence of resistant bacterial strains along with poor healthcare standards.39 Moreover, data to monitor trends and susceptibility patterns against antibiotics are infrequently produced.4,13 Hence, the implementation of strong antibiotic stewardship programs is crucial to track the spread of MDR pathogens in Bangladesh. As the problem of AMR is multi-sectoral, a multi-disciplinary and holistic approach is necessary to ensure the best use of antibiotics, prescribe the right antibiotics, and preserve the ‘last resort’ antibiotics.1,39 The overall findings of the present study will provide useful information to the local healthcare authorities towards the development of a strong antimicrobial stewardship program.
The major limitation of this study was that the clinical samples were collected from the highly urbanized Dhaka region only, which limited the generalizability of our results. For tracking the resistance patterns of Salmonella spp., a more detailed study with a larger sample size, covering both rural and urban areas, is recommended. More facilities providing affordable microbiological tests should be provided nationwide to enable physicians to optimize antibiotic prescriptions. Simultaneously, the prohibition against dispensing antibiotics without a prescription should be strictly enforced and monitored.
Finally, identifying the constantly evolving genes responsible for resistance to pathogens is essential. Continuous genetic research is essential to stay a step ahead of the pathogens. Several past studies have revealed intriguing commonalities in the resistance trends of pathogens in the South Asian region. This is an opportunity for strong regional cooperation regarding genetic research on resistant pathogens. The altruistic cooperation among the international healthcare industry during the COVID-19 pandemic has shown us the way. Continuing in the same spirit, we can turn the tide against the formidable challenge of antibiotic resistance.
Conclusion
The current study reveals a significant and growing prevalence of MDR Salmonella isolates in Dhaka, Bangladesh. This calls for urgent and effective measures to monitor and regulate the prescribing, sales, and consumption of antibiotics. Countrywide expansion of screening facilities for microbial infections and causative agents using standardized approaches is highly desirable. It is crucial to conduct systematic and focused studies on the prevalence and incidence of antibiotic resistance including MDR and XDR patterns. We also recommend stronger cooperation between South Asian countries to initiate joint research on the genetic aspects of antibiotic resistance, and to reach targeted remedies.
Disclosure
The authors declared no conflicts of interest. No funding was received for this study.
Acknowledgments
We thank the North South University, Bangladesh, for providing facilities to carry out this study.
references
- 1. Asokan GV, Ramadhan T, Ahmed E, Sanad H. WHO global priority pathogens list: a bibliometric analysis of Medline-PubMed for knowledge mobilization to infection prevention and control practices in Bahrain. Oman Med J 2019 May;34(3):184-193.
- 2. Sarker S, Mahmud S, Sultana R, Biswas R, Sarkar PP, Munayem MA, et al. Quality assessment of surface and drinking water of Nakla Paurosova, Sherpur, Bangladesh. Adv Microbiol 2019 Aug;9(8):703-727 .
- 3. Al-Yamani A, Khamis F, Al-Zakwani I, Al-Noomani H, Al-Noomani J, Al-Abri S. Patterns of antimicrobial prescribing in a tertiary care hospital in Oman. Oman Med J 2016 Jan;31(1):35-39.
- 4. Ahmed I, Rabbi MB, Sultana S. Antibiotic resistance in Bangladesh: A systematic review. Int J Infect Dis 2019 Mar;80:54-61.
- 5. Popa GL, Papa MI. Salmonella spp. infection - a continuous threat worldwide. Germs 2021 Mar;11(1):88-96.
- 6. Ketema L, Ketema Z, Kiflu B, Alemayehu H, Terefe Y, Ibrahim M, et al. Prevalence and antimicrobial susceptibility profile of Salmonella serovars isolated from slaughtered cattle in Addis Ababa, Ethiopia. Biomed Res Int 2018 Nov;2018(1):9794869.
- 7. Khan AI. Enteric disease surveillance in Bangladesh: findings from a nationwide hospital-based surveillance in ten selected sites. National Bulletin of Public Health (NBPH); 2018.
- 8. Cho S, Hiott LM, House SL, Woodley TA, McMillan EA, Sharma P, et al. Analysis of salmonella enterica isolated from a mixed-use watershed in Georgia, USA: antimicrobial resistance, serotype diversity, and genetic relatedness to human isolates. Appl Environ Microbiol 2022 May;88(10):e0039322.
- 9. Jaja IF, Bhembe NL, Green E, Oguttu J, Muchenje V. Molecular characterisation of antibiotic-resistant Salmonella enterica isolates recovered from meat in South Africa. Acta Trop 2019 Feb;190:129-136.
- 10. Marzan M, Islam DZ, Lugova H, Krishnapillai A, Haque M, Islam S. Knowledge, attitudes, and practices of antimicrobial uses and resistance among public university students in Bangladesh. Infect Drug Resist 2021 Feb;14:519-533.
- 11. Bell AG, Thornber K, Chaput DL, Hasan NA, Alam MM, Haque MM, et al. The diversity and ubiquity of antibiotic resistance genes in finfish culture ponds in Bangladesh. BioRxiv 2022 Sep 14:2022-09 .
- 12. Qayyum S, Basharat S, Mian AH, Qayum S, Ali M, Changsheng P, et al. Isolation, identification and antibacterial study of pigmented bacteria. Appl Nanosci 2020 Dec;10:4495-4503 .
- 13. Jain P, Bepari AK, Sen PK, Rafe T, Imtiaz R, Hossain M, et al. High prevalence of multiple antibiotic resistance in clinical E. coli isolates from Bangladesh and prediction of molecular resistance determinants using WGS of an XDR isolate. Sci Rep 2021 Nov;11(1):22859.
- 14. Benkova M, Soukup O, Marek J. Antimicrobial susceptibility testing: currently used methods and devices and the near future in clinical practice. J Appl Microbiol 2020 Oct;129(4):806-822.
- 15. Bebe T, Odetoyin B, Bolarinwa R. Occurrence of multidrug-resistant uropathogens implicated in asymptomatic bacteriuria in adults with sickle cell disease in ile-ife, Southwest Nigeria. Oman Med J 2020 Mar;35(2):e109.
- 16. Ballash GA, Albers AL, Mollenkopf DF, Sechrist E, Adams RJ, Wittum TE. Antimicrobial resistant bacteria recovered from retail ground meat products in the US include a Raoultella ornithinolytica co-harboring blaKPC-2 and blaNDM-5. Sci Rep 2021 Jul;11(1):14041.
- 17. Ali Shah SA, Nadeem M, Syed SA, Fatima Abidi ST, Khan N, Bano N. Antimicrobial sensitivity pattern of salmonella typhi: emergence of resistant strains. Cureus 2020 Nov;12(11):e11778.
- 18. Al Saifi SA, Al Adawi B, Burney I. Patterns of bacterial isolates and their resistance to antibiotics in patients with chemotherapy-induced febrile neutropenia at a university hospital. Oman Med J 2021 Jul;36(4):e290.
- 19. Uddin MS, Hoq MI, Ali MS, Rahman MM, Islam KS. Antibiotic resistance pattern of salmonella spp. isolated from stool samples of hospitalized diarrheal patients in Bangladesh. Asian J. Med. Biol 2018;3(4):534-538 .
- 20. Soltan Dallal MM, Khalilian M, Masoumi Asl H, Bakhtiari R, Davoodabadi A, Sharifi Yazdi MK, et al. Molecular epidemiology and antimicrobial resistance of salmonella spp. isolated from resident patients in Mazandaran Province, Northern Iran. J Food Qual Hazards Control 2016 Dec;3(4):146-151.
- 21. Arabestani MR, Rastiyani S, Alikhani MY, Mousavi SF. The relationship between prevalence of antibiotics resistance and virulence factors genes of MRSA and MSSA strains isolated from clinical samples. Oman Med J 2018 Mar;33(2):134-140.
- 22. El-Tayeb MA, Ibrahim AS, Al-Salamah AA, Almaary KS, Elbadawi YB. Prevalence, serotyping and antimicrobials resistance mechanism of Salmonella enterica isolated from clinical and environmental samples in Saudi Arabia. Braz J Microbiol 2017;48(3):499-508.
- 23. Dembélé R, Konaté A, Soulama I, Kagambèga A, Kaboré WA, Cissé H. Prevalence of multidrug-resistant salmonella enterica and associated factors among under five children with diarrhea in rural Burkina Faso. Clin Biotechnol Microbiol. 2018;3(1):566-576.
- 24. Mbuthia SW, Some ES, Suleiman M, Mbuthia OW, Ngayo MO. Assessment of clinical presentation, performance of diagnostic methods and antibiotic susceptibility testing for salmonella among patients attending Kangema sub-county hospital, Kenya. Open J Epidemiol 2022 Oct;12(4):449-469.
- 25. Zahid I, Sarwar A, Hussain A, Sohail M, Amin A. Antibiotyping and genotyping of extensively drug-resistant (XDR) Salmonella sp. isolated from clinical samples of Lahore, Pakistan. J Appl Microbiol 2022 Jan;132(1):633-641.
- 26. Islam K, Mahmud R, Chowdhury MK, Hossain FS, Biswas PK, Sarker S. Recent sensitivity pattern of salmonella typhi in a private hospital. J Med 2018;19(1):15.
- 27. Rao PS, Rajashekar V, Varghese GK, Shivananda PG. Emergence of multidrug-resistant Salmonella typhi in rural southern India. Am J Trop Med Hyg 1993 Jan;48(1):108-111.
- 28. Qamar FN, Azmatullah A, Kazi AM, Khan E, Zaidi AK. A three-year review of antimicrobial resistance of Salmonella enterica serovars Typhi and Paratyphi A in Pakistan. J Infect Dev Ctries 2014 Aug;8(8):981-986.
- 29. Andrews JR, Qamar FN, Charles RC, Ryan ET. Extensively drug-resistant typhoid—are conjugate vaccines arriving just in time? N Engl J Med 2018 Oct;379(16):1493-1495.
- 30. Katiyar A, Sharma P, Dahiya S, Singh H, Kapil A, Kaur P. Genomic profiling of antimicrobial resistance genes in clinical isolates of Salmonella Typhi from patients infected with Typhoid fever in India. Sci Rep 2020 May;10(1):8299.
- 31. Mannan A, Shohel M, Rajia S, Mahmud NU, Kabir S, Hasan I. A cross sectional study on antibiotic resistance pattern of Salmonella typhi clinical isolates from Bangladesh. Asian Pac J Trop Biomed 2014 Apr;4(4):306-311.
- 32. Habib ZH, Golam Rasul SB, Alam MA, Alam MA, Bably NN, Khan IA, et al. The findings of antimicrobial resistance surveillance in Bangladesh (2016-2020). medRxiv 2021 Jun 14:2021-06 .
- 33. Wong MH, Yan M, Chan EW, Biao K, Chen S. Emergence of clinical Salmonella enterica serovar Typhimurium isolates with concurrent resistance to ciprofloxacin, ceftriaxone, and azithromycin. Antimicrob Agents Chemother 2014 Jul;58(7):3752-3756.
- 34. Abdullah FE, Tanvir SB, Syed AZ, Haq MF, Shariq A, Ahmed S. Prevalence of salmonella typhi and paratyphi in urine cultures and their antibiotic sensitivity patterns in Karachi. European J Biotechnol Biosc 2015 Mar;3(3):35-38.
- 35. Qamar FN, Yousafzai MT, Dehraj IF, Shakoor S, Irfan S, Hotwani A, et al. Antimicrobial resistance in typhoidal salmonella: surveillance for enteric fever in Asia project, 2016–2019. Clin Infect Dis 2020 Dec;71(Suppl 3):S276-S284.
- 36. Klemm EJ, Shakoor S, Page AJ, Qamar FN, Judge K, Saeed DK, et al. Emergence of an extensively drug-resistant salmonella enterica serovar Typhi clone harboring a promiscuous plasmid encoding resistance to fluoroquinolones and third-generation cephalosporins. mBio 2018 Feb;9(1):10-128.
- 37. Ambreen A, Batool A, Ahmad Q, Nisar A, Rasool MH, Qamar MU. Emergence of MDR and XDR salmonella enterica serovar typhi from pediatric septicemia patients in Lahore. Int J Infect Dis 2020 Dec;101:443 .
- 38. Abd El-Aziz NK, Tartor YH, Gharieb RM, Erfan AM, Khalifa E, Said MA, et al. Extensive drug-resistant salmonella enterica isolated from poultry and humans: prevalence and molecular determinants behind the co-resistance to ciprofloxacin and tigecycline. Front Microbiol 2021 Nov;12:738784.
- 39. Hoque R, Ahmed SM, Naher N, Islam MA, Rousham EK, Islam BZ, et al. Tackling antimicrobial resistance in Bangladesh: A scoping review of policy and practice in human, animal and environment sectors. PLoS One 2020 Jan;15(1):e0227947.